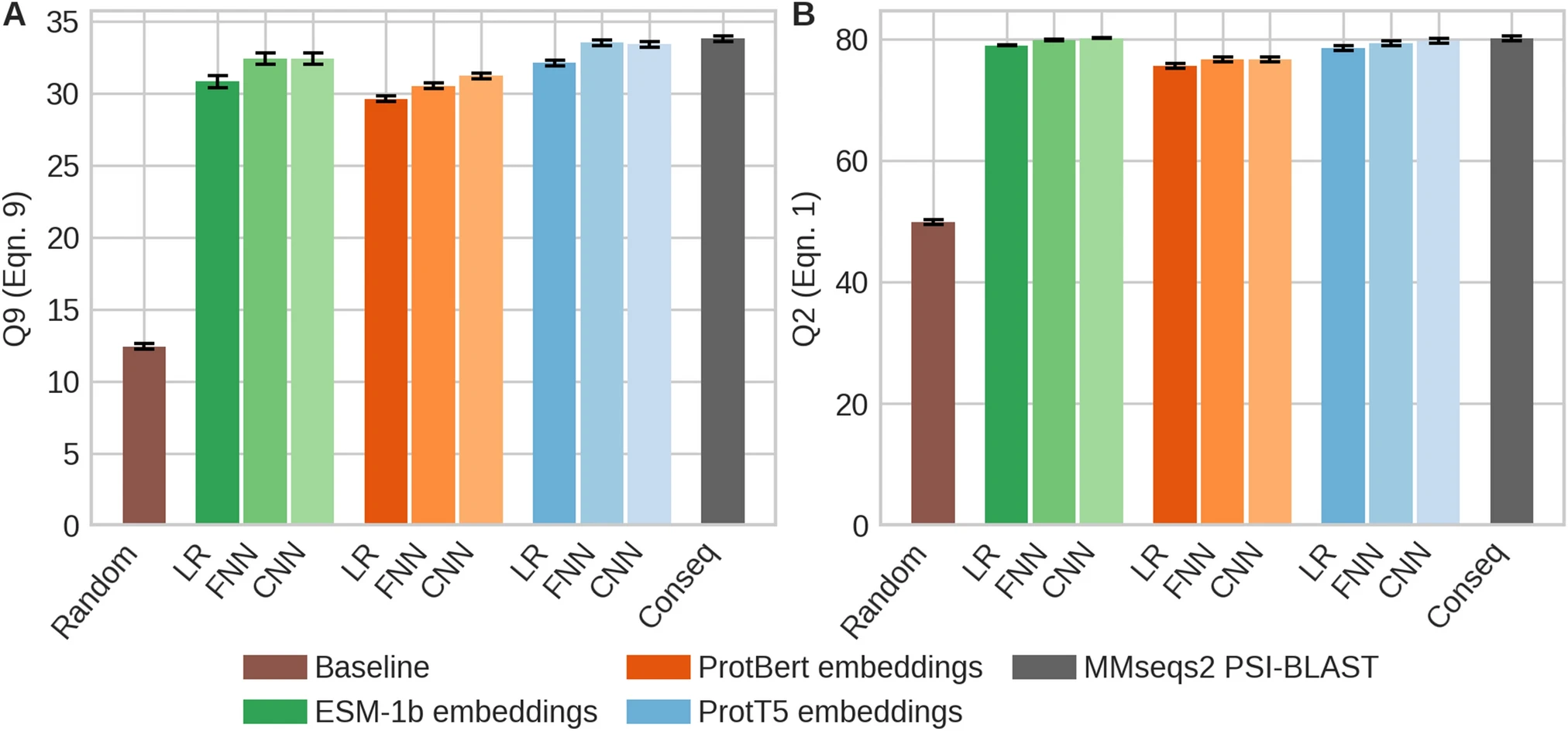
The dataset provided here aims at predicting protein conservation (9-state).
The possible conservation states for each residue (assigned to TARGET) range from 1 to 9 (1 = very variable, 9 =
very conserved).
The provided dataset is compiled as follows:
- Training / Validation / Test: FLIP-sampled
Removed Sequences
There were a couple of sequences in the FLIP that were missing a label. These were removed from the dataset:- 1unq-A
- 5xj5-A
- 1hlb-A
- 2aal-A
- 6ndt-B
- 2gud-A
- 3vla-A
- 1jly-A
- 5ima-A
- 1kqf-A
- 3l8w-A
- 5vf5-A
- 6a56-A
- 1xkp-A
- 2gnx-A
- 3ar4-A
- 3w19-C
- 2xsk-A
- 5m29-A
- 5o2x-A
- 5vht-A
- 5y0t-A
- 4fzv-B
- 4pio-A
- 5xdh-A
- 6hik-L
- 4kbx-A
- 2ofc-A
- 4zbl-A
- 5ec6-A
- 3u5s-A
- 1pya-B
- 2cnq-A
- 1nth-A
- 5e5y-A
- 5jiw-A
- 1n13-B
- 3esm-A
- 2omk-A
- 4mt2-A
- 5a7v-A
The dataset is provided in biotrainer-ready fasta format. Each entry contains a sequence and a header, providing the sequence id, the set (train/val/test) and the target label.
Marquet et al. (2021) provide the following benchmarks for this dataset:
@article{marquet2021embeddings,
title={Embeddings from protein language models predict conservation and variant effects},
author={Marquet, C{\'e}line and Heinzinger, Michael and Olenyi, Tobias and Dallago, Christian and Erckert, Kyra and Bernhofer, Michael and Nechaev, Dmitrii and Rost, Burkhard},
journal={Human genetics},
pages={1--19},
year={2021},
abstract = "{The emergence of SARS-CoV-2 variants stressed the demand for tools allowing to interpret the effect of single amino acid variants (SAVs) on protein function. While Deep Mutational Scanning (DMS) sets continue to expand our understanding of the mutational landscape of single proteins, the results continue to challenge analyses. Protein Language Models (pLMs) use the latest deep learning (DL) algorithms to leverage growing databases of protein sequences. These methods learn to predict missing or masked amino acids from the context of entire sequence regions. Here, we used pLM representations (embeddings) to predict sequence conservation and SAV effects without multiple sequence alignments (MSAs). Embeddings alone predicted residue conservation almost as accurately from single sequences as ConSeq using MSAs (two-state Matthews Correlation Coefficient—MCC—for ProtT5 embeddings of 0.596 ± 0.006 vs. 0.608 ± 0.006 for ConSeq). Inputting the conservation prediction along with BLOSUM62 substitution scores and pLM mask reconstruction probabilities into a simplistic logistic regression (LR) ensemble for Variant Effect Score Prediction without Alignments (VESPA) predicted SAV effect magnitude without any optimization on DMS data. Comparing predictions for a standard set of 39 DMS experiments to other methods (incl. ESM-1v, DeepSequence, and GEMME) revealed our approach as competitive with the state-of-the-art (SOTA) methods using MSA input. No method outperformed all others, neither consistently nor statistically significantly, independently of the performance measure applied (Spearman and Pearson correlation). Finally, we investigated binary effect predictions on DMS experiments for four human proteins. Overall, embedding-based methods have become competitive with methods relying on MSAs for SAV effect prediction at a fraction of the costs in computing/energy. Our method predicted SAV effects for the entire human proteome (~ 20 k proteins) within 40 min on one Nvidia Quadro RTX 8000. All methods and data sets are freely available for local and online execution through bioembeddings.com, https://github.com/Rostlab/VESPA, and PredictProtein.}"
publisher={Springer}
}@InProceedings{dallago2021flip,
author = {Christian Dallago and Jody Mou and Kadina E Johnston and Bruce Wittmann and Nick Bhattacharya and Samuel Goldman and Ali Madani and Kevin K Yang},
booktitle = {Thirty-fifth Conference on Neural Information Processing Systems Datasets and Benchmarks Track (Round 2)},
title = {{FLIP}: Benchmark tasks in fitness landscape inference for proteins},
year = {2021},
url = {https://openreview.net/forum?id=p2dMLEwL8tF},
}The RAW data downloaded from the aforementioned publications is subject to Attribution 4.0 International (CC BY 4.0). Modified data available in this repository falls under AFL-3.
This is an Open Access article distributed under the terms of the Creative Commons Attribution License (https://creativecommons.org/licenses/by/4.0/), which permits unrestricted reuse, distribution, and reproduction in any medium, provided the original work is properly cited.