-
Notifications
You must be signed in to change notification settings - Fork 24
Web Application
A web app that is launched from the command line can be used to view and analyse results from a set of predictions that you have made. This is an improved and much easier to use form of a previous web interface called epitopemap and replaces it.
Note: this app is still under early development, suggestions for additional functionality are welcome.
After you have made some predictions you can run the app using epitopepredict -s. This opens a new web page in your browser. You then enter the path (folder) where your results were saved in the form and press submit. This should refresh your form with a drop down list of all the available sequences/proteins.
There are several ways to view a set of binding predictions, all of which allow views for whichever predictors you have used. There is currently
Summarizes the results for multiple sequences in one page. You can choose to view the table of all predicted binders, promiscuous binders or a summary over each sequence. These tables can be downloaded to csv files.
For viewing the detailed results for a single sequence, often representing a protein coding sequence. Graphical views of the prediction scoring across the sequence are designed to provide a quick look at the pattern of peptide binding prediction in multiple alleles. By default a track view of each allele/predictor is shown as below:
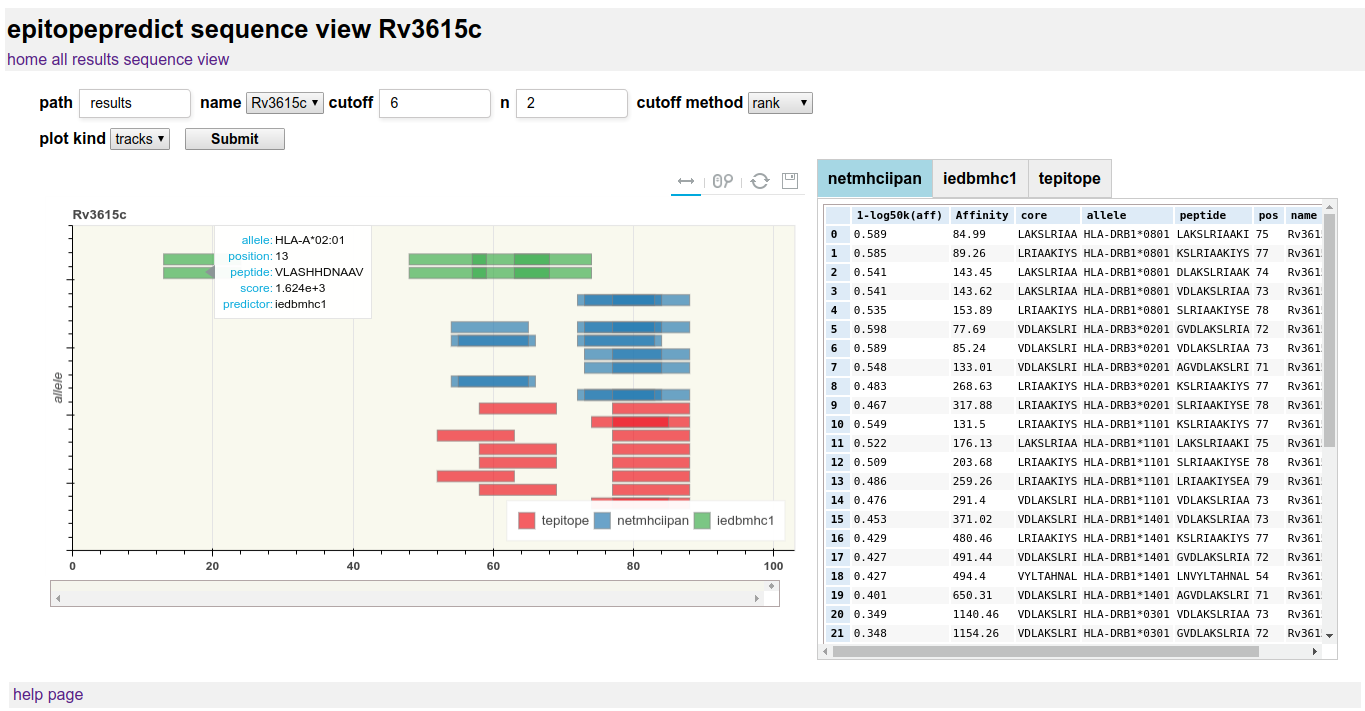
Track view is useful for overall results over protein-length and longer sequences. The plot can be zoomed and panned using the mouse. A hover tooltip shows the particular peptide details.
- Improved graphical features for genome based prediction.
- Location of clusters of binders in sequences.
- Export of peptide lists/n-mers for experimental use.
- Mutation/conservation analysis.
- Edit config and run predictions from web page.