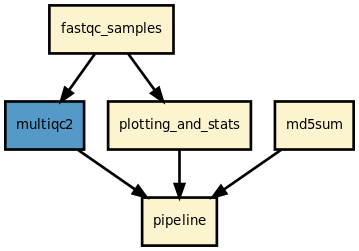
This is the fastqc pipeline from the Sequana projet
| Overview: | Runs fastqc (or falco) and multiqc on a set of sequencing data to produce quality control reports |
|---|---|
| Input: | A set of FastQ files (paired or single-end), compressed or not |
| Output: | An HTML summary report with individual FastQC reports and a multi-sample MultiQC report |
| Status: | Production |
| Documentation: | This README file and https://sequana.readthedocs.io |
| Citation: | Cokelaer et al, (2017), 'Sequana': a Set of Snakemake NGS pipelines, Journal of Open Source Software, 2(16), 352, JOSS DOI https://doi:10.21105/joss.00352 |
If you already have all requirements, you can install the package using pip:
pip install sequana_fastqc --upgrade
You will need third-party software such as fastqc or falco. Please see below for details.
Scan FastQ files in a directory and set up the pipeline (replace DATAPATH with your input directory):
sequana_fastqc --input-directory DATAPATH
To use falco instead of fastqc:
sequana_fastqc --input-directory DATAPATH --method falco
To skip the MultiQC report (useful when memory is limited):
sequana_fastqc --input-directory DATAPATH --skip-multiqc
This creates a fastqc/ directory with the pipeline and configuration file. Execute the pipeline locally:
cd fastqc sh fastqc.sh
If you are familiar with Snakemake, you can also run the pipeline directly:
snakemake -s fastqc.rules --cores 4 --stats stats.txt
See .sequana/profile/config.yaml to tune Snakemake behaviour (cores, cluster settings, etc.).
With apptainer, initiate the working directory as follows:
sequana_fastqc --input-directory DATAPATH --use-apptainer
Images are downloaded in the working directory but you can store them in a shared location:
sequana_fastqc --input-directory DATAPATH --use-apptainer --apptainer-prefix ~/.sequana/apptainers
and then:
cd fastqc sh fastqc.sh
This pipeline requires the following executables (install via bioconda/conda):
- fastqc — quality control tool for sequencing data (default)
- falco — faster drop-in replacement for fastqc (optional,
--method falco) - multiqc — aggregated HTML report across samples
Install all dependencies at once:
mamba env create -f environment.yml
This pipeline runs fastqc (or falco) in parallel on the input FastQ files, then aggregates results with MultiQC. A sequana summary report is also produced with per-sample statistics and quality plots.
QC method (--method):
fastqc(default) — standard FastQC; handles FastQ, BAM, and SAM inputsfalco— faster alternative; FastQ inputs only
Optional outputs:
- MultiQC report (
multiqc/multiqc_report.html) — enabled by default, disable with--skip-multiqc - MD5 checksums of all input files (
md5.txt) - Tree browser of individual FastQC HTML reports (
tree.html)
This pipeline has been tested on several hundreds of MiSeq, NextSeq, MiniSeq, ISeq100, and PacBio runs.
Here is the latest documented configuration file to be used with the pipeline. Each rule used in the pipeline may have a section in the configuration file.
| Version | Description |
|---|---|
| 1.10.0 |
|
| 1.9.0 |
|
| 1.8.2 |
|
| 1.8.1 |
|
| 1.8.0 |
|
| 1.7.1 |
|
| 1.7.0 |
|
| 1.6.2 |
|
| 1.6.1 |
|
| 1.6.0 |
|
| 1.5.0 |
|
| 1.4.2 |
|
| 1.4.1 |
|
| 1.4.0 |
|
| 1.3.0 |
|
| 1.2.0 |
|
| 1.1.0 |
|
| 1.0.1 |
|
| 1.0.0 |
|
| 0.9.15 |
|
| 0.9.14 |
|
| 0.9.13 |
|
| 0.9.12 |
|
| 0.9.11 |
|
| 0.9.10 |
|
| 0.9.9 |
|
| 0.9.8 |
|
| 0.9.7 |
|
| 0.9.6 | add the readtag option |
To contribute to this project, please take a look at the Contributing Guidelines first. Please note that this project is released with a Code of Conduct. By contributing to this project, you agree to abide by its terms.