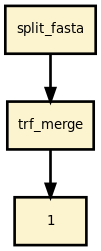
This is the trf (tandem repeat finder) pipeline from the Sequana project
| Overview: | run TRF on several large datasets |
|---|---|
| Input: | a set of FastA files |
| Output: | TRF output + images |
| Status: | draft |
| Citation: | Cokelaer et al, (2017), ‘Sequana’: a Set of Snakemake NGS pipelines, Journal of Open Source Software, 2(16), 352, JOSS DOI doi:10.21105/joss.00352 |
If you already have all requirements, you can install the packages using pip:
pip install sequana_trf --upgrade
Otherwise, you can create a sequana_trf conda environment executing:
conda env create -f environment.yml
and later activate the environment:
conda activate sequana_trf
A third option is to install the pipeline with pip method (see above) and use singularity as explained afterwards.
sequana_trf --help sequana_trf --input-directory DATAPATH
This creates a directory with the pipeline and configuration file. You will then need to execute the pipeline:
cd trf sh trf.sh # for a local run
This launch a snakemake pipeline. If you are familiar with snakemake, you can retrieve the pipeline itself and its configuration files and then execute the pipeline yourself with specific parameters:
snakemake -s trf.rules -c config.yaml --cores 4 --stats stats.txt
Or use sequanix interface.
This pipeline requires the following executable(s):
- trf
This pipeline runs trf in parallel on the input fastq files (paired or not). A brief sequana summary report is also produced.
Here is the latest documented configuration file to be used with the pipeline. Each rule used in the pipeline may have a section in the configuration file.
| Version | Description |
|---|---|
| 1.0.0 |
|
| 0.10.0 |
|
| 0.9.3 |
|
| 0.9.2 |
|
| 0.9.0 | First release. |